 I’ve been having discussions with my colleague, Alex Kandel, about a software tool he’s been working on. He has 20,000 or so STM images that his group has taken over the past five years, and he is building a web tool that will let his group search the images based on criteria like tip-surface voltage bias, surface preparation, scan rate, who took the images, etc.
I’ve been having discussions with my colleague, Alex Kandel, about a software tool he’s been working on. He has 20,000 or so STM images that his group has taken over the past five years, and he is building a web tool that will let his group search the images based on criteria like tip-surface voltage bias, surface preparation, scan rate, who took the images, etc.
So far, he’s been using Zoph, which is a photo gallery which has a nice search tool for the EXIF and IPTC data in JPEG and TIFF images. Zoph is a nice tool, but given the difficulty of writing EXIF headers into images with the common tools (ImageMagick, gd, netpbm), he’s had to hijack the image import functions and read the “extra” data from text files created by his microscopes.
EXIF and IPTC are great examples of file formats which store meta-data in the same file with the primary data. It makes a lot of sense to store the experimental parameters that generated an image in with the image itself.
My favorite gallery software, Gallery2 lets you view, but not search the EXIF data. Zoph‘s search functionality is a lot more extensive, so it was the natural choice for Alex’s tool.
There are many of areas of science (scanning probe microscopy, optical microscopy, various forms of astronomy) which use image data as their primary source of quantitative information, and there are some really wonderful open source image gallery tools that have been written to organize home and professional photography. Just a few more pieces are necessary to make these powerful scientific tools as well:

- A unix command-line tool or library that gives us the ability to insert and edit EXIF/IPTC data in arbitrary images.
- Data import modules for Zoph or Gallery2 that use the EXIF/IPTC data to populate the database with details from the metadata stored in the image file.
- Extensible search modules for Zoph or Gallery2 that make it trivial to search arbitrary field names in this data.
These would turn good amateur photography tools into powerful scientific image managers.
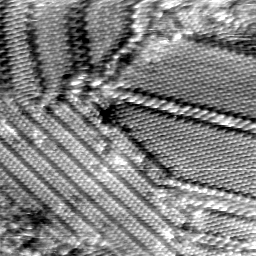
Update: Alex found Exiv2 which can read and write EXIF and IPTC data directly. The second image above is a sample STM image which has a few “interesting” EXIF fields:
% exiv2 -pt 09090400BT.jpg
Exif.Image.DocumentName Ascii 15 09090400BT.SM3
Exif.Image.XResolution SLong 1 400
Exif.Image.YResolution SLong 1 400
Exif.Image.PlanarConfiguration Short 1 0
Exif.Image.ExifTag Long 1 89
Exif.Photo.ExposureTime SLong 1 23 s
Exif.Photo.SpectralSensitivity Ascii 16 lowpass filter
Exif.Photo.DateTimeOriginal Ascii 21 2004:09:09 14:27:26Exif.Photo.BrightnessValue SRational 1 759
Exif.Photo.ExposureBiasValue SRational 1 +249
Exif.Photo.Flash Short 1 Yes
Exif.Photo.UserComment Undefined 80 Octanethiol SAM first imaged on 9-8-04, left in pink thiol box overnight
Although Exiv2 looks like it might be the key, Alex notes a few problems remaining with this approach:
- He’s saving all values as ints or rationals because EXIF doesn’t seem to support floats.
- He’s storing data in some infrequently used EXIF fields, and not all parsers will read it by default.
[tags]science, images, metadata[/tags]
Have you seen Fly Express? It’s limited to Drosophila embryos, but it’s a searchable database. Pretty cool. I think the search algorithm is based on those implemented in face identification software.
Hi,
beside exiv2, there is ExifTool as well, I don’t know if it resolves the problem you have found with exiv2.